Lineage Workflow
Learn how to configure the Lineage workflow from the UI to ingest Lineage data from your data sources.
Checkout the documentation of the connector you are using to know if it supports automated lineage workflow.
If your database service is not yet supported, you can use this same workflow by providing a Query Log file!
Learn how to do so 👇
UI Configuration
Once the metadata ingestion runs correctly and we are able to explore the service Entities, we can add Entity Lineage information.
This will populate the Lineage tab from the Table Entity Page.
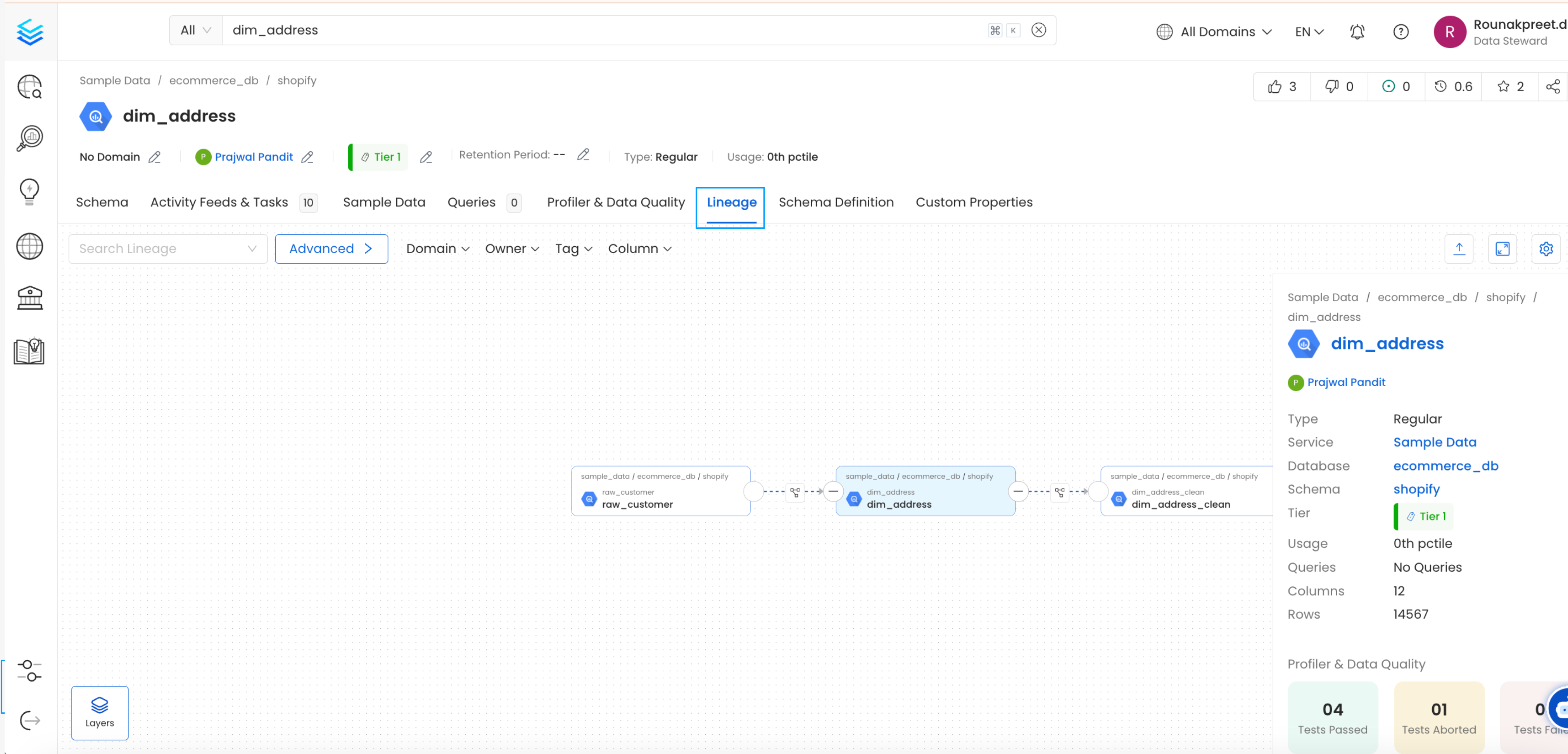
Table Entity Page
We can create a workflow that will obtain the query log and table creation information from the underlying database and feed it to OpenMetadata. The Lineage Ingestion will be in charge of obtaining this data.
1. Add a Lineage Ingestion
From the Service Page, go to the Ingestions tab to add a new ingestion and click on Add Lineage Ingestion.
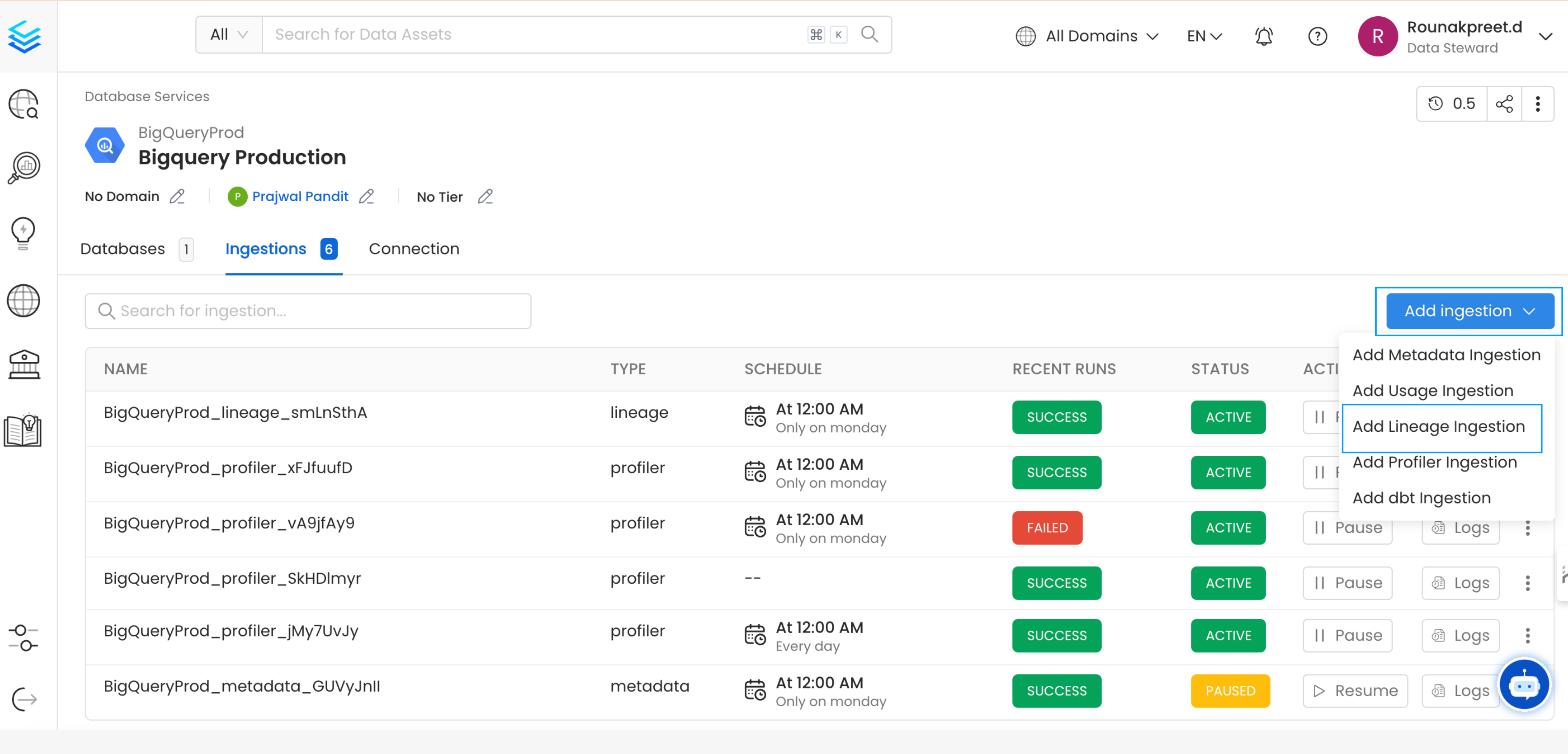
Add Ingestion
2. Configure the Lineage Ingestion
Here you can enter the Lineage Ingestion details:
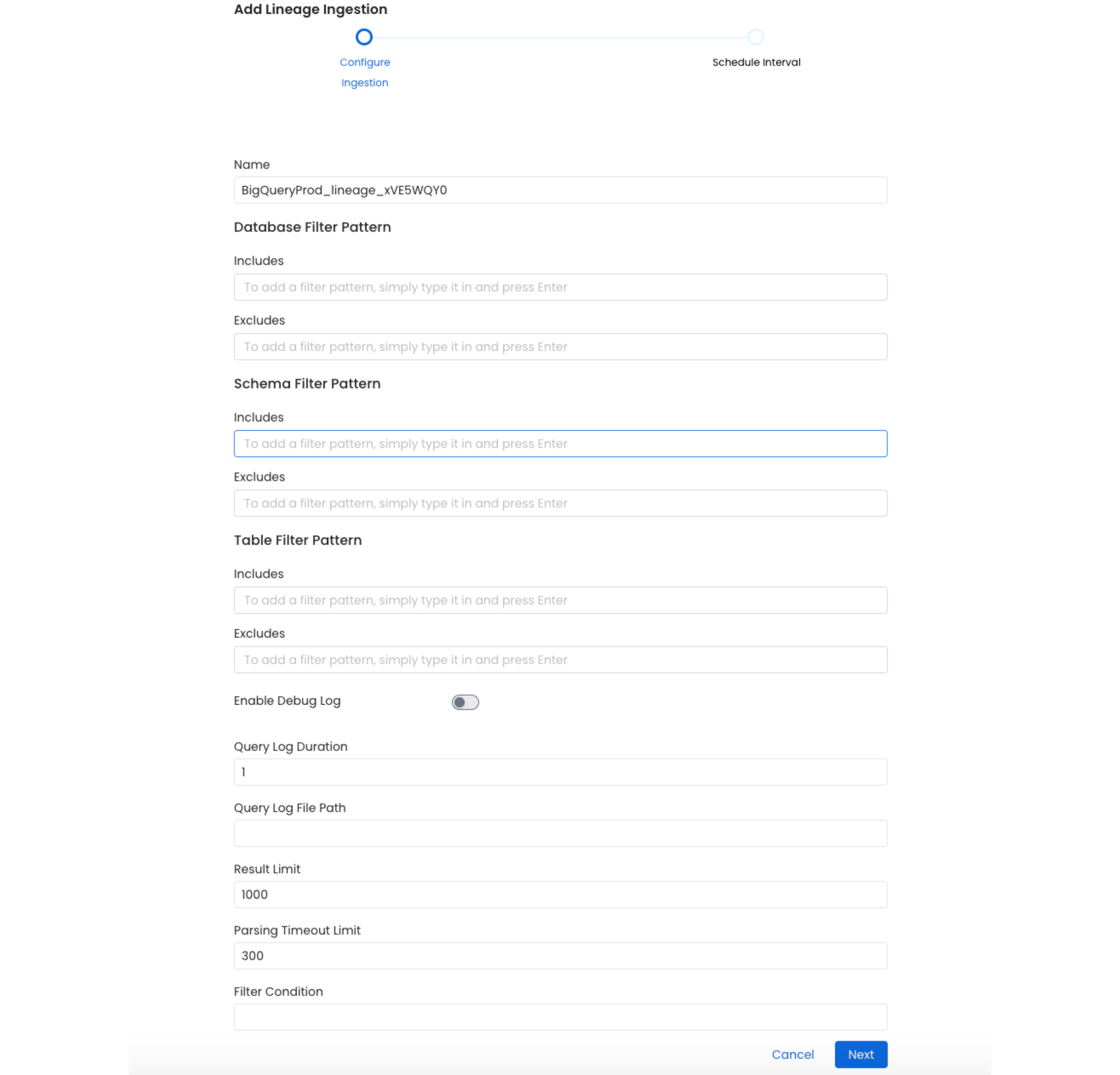
Configure the Lineage Ingestion
Lineage Options
Query Log Duration
Specify the duration in days for which the lineage should capture lineage data from the query logs. For example, if you specify 2 as the value for the duration, the data lineage will capture lineage information for 48 hours prior to when the ingestion workflow is run.
Result Limit
Set the limit for the query log results to be run at a time.
3. Schedule and Deploy
After clicking Next, you will be redirected to the Scheduling form. This will be the same as the Metadata Ingestion. Select your desired schedule and click on Deploy to find the lineage pipeline being added to the Service Ingestions.
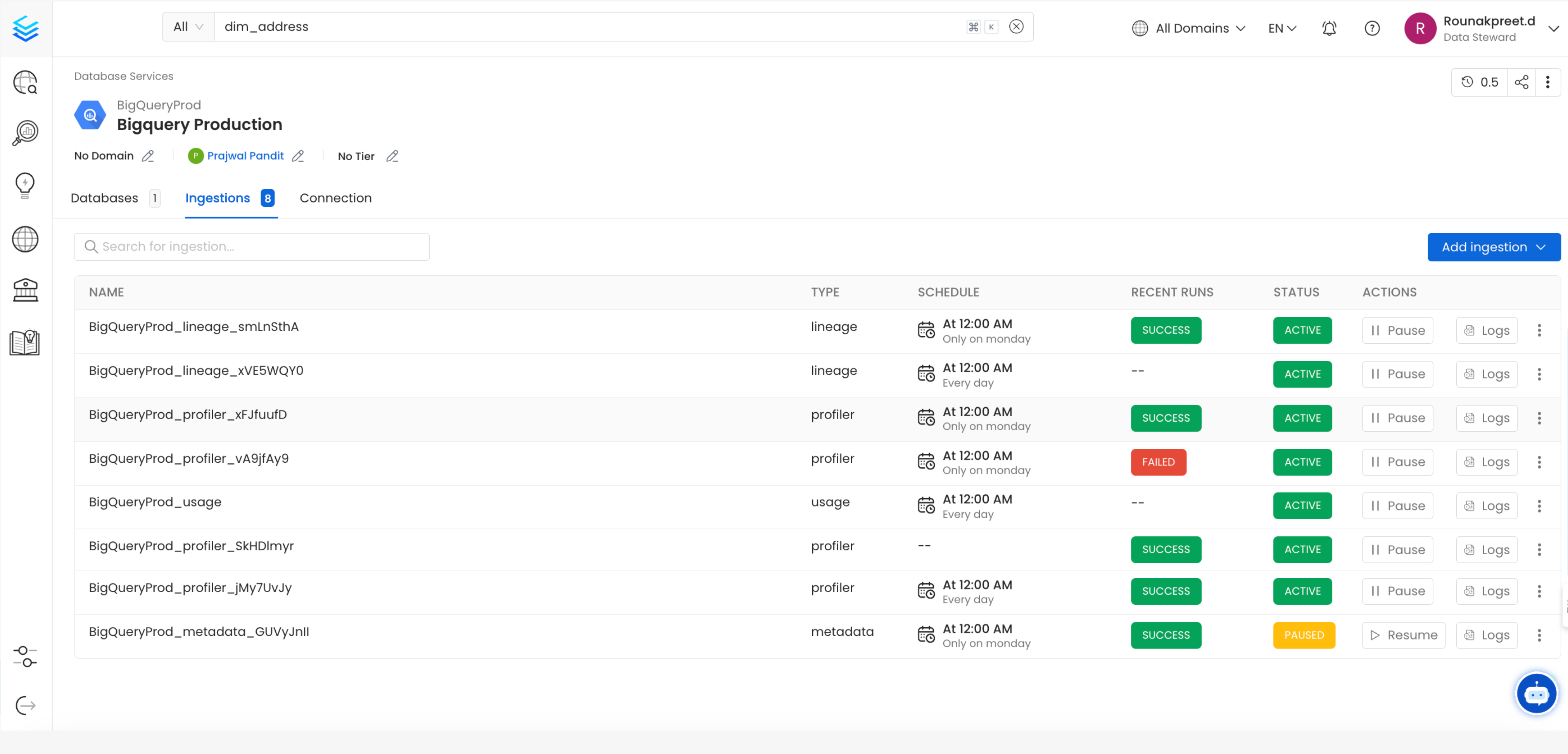
View Service Ingestion pipelines
YAML Configuration
In the connectors section we showcase how to run the metadata ingestion from a JSON/YAML file using the Airflow SDK or the CLI via metadata ingest. Running a lineage workflow is also possible using a JSON/YAML configuration file.
This is a good option if you wish to execute your workflow via the Airflow SDK or using the CLI; if you use the CLI a lineage workflow can be triggered with the command metadata ingest -c FILENAME.yaml. The serviceConnection config will be specific to your connector (you can find more information in the connectors section), though the sourceConfig for the lineage will be similar across all connectors.
Lineage
After running a Metadata Ingestion workflow, we can run Lineage workflow. While the serviceName will be the same to that was used in Metadata Ingestion, so the ingestion bot can get the serviceConnection details from the server.
1. Define the YAML Config
This is a sample config for BigQuery Lineage:
Source Configuration - Source Config
You can find all the definitions and types for the sourceConfig here.
queryLogDuration: Configuration to tune how far we want to look back in query logs to process lineage data in days.
parsingTimeoutLimit: Configuration to set the timeout for parsing the query in seconds.
filterCondition: Condition to filter the query history.
resultLimit: Configuration to set the limit for query logs.
queryLogFilePath: Configuration to set the file path for query logs.
databaseFilterPattern: Regex to only fetch databases that matches the pattern.
schemaFilterPattern: Regex to only fetch tables or databases that matches the pattern.
tableFilterPattern: Regex to only fetch tables or databases that matches the pattern.
overrideViewLineage: Set the 'Override View Lineage' toggle to control whether to override the existing view lineage.
processViewLineage: Set the 'Process View Lineage' toggle to control whether to process view lineage.
processQueryLineage: Set the 'Process Query Lineage' toggle to control whether to process query lineage.
processStoredProcedureLineage: Set the 'Process Stored ProcedureLog Lineage' toggle to control whether to process stored procedure lineage.
threads: Number of Threads to use in order to parallelize lineage ingestion.
- You can learn more about how to configure and run the Lineage Workflow to extract Lineage data from here
2. Run with the CLI
After saving the YAML config, we will run the command the same way we did for the metadata ingestion: