Metadata Ingestion
Connection Details
Connection Details
- username: Enter the username of your Amundsen user in the Username field. The specified user should be authorized to read all databases you want to include in the metadata ingestion workflow.
- password: Enter the password for your amundsen user in the Password field.
- hostPort: Host and port of the Amundsen Neo4j Connection. This expect a URI format like: bolt://localhost:7687.
- maxConnectionLifeTime (optional): Maximum connection lifetime for the Amundsen Neo4j Connection
- validateSSL (optional): Enable SSL validation for the Amundsen Neo4j Connection.
- encrypted (Optional): Enable encryption for the Amundsen Neo4j Connection.
Test the Connection
Once the credentials have been added, click on Test Connection and Save the changes.

Configure Metadata Ingestion
In this step we will configure the metadata ingestion pipeline,
Please follow the instructions below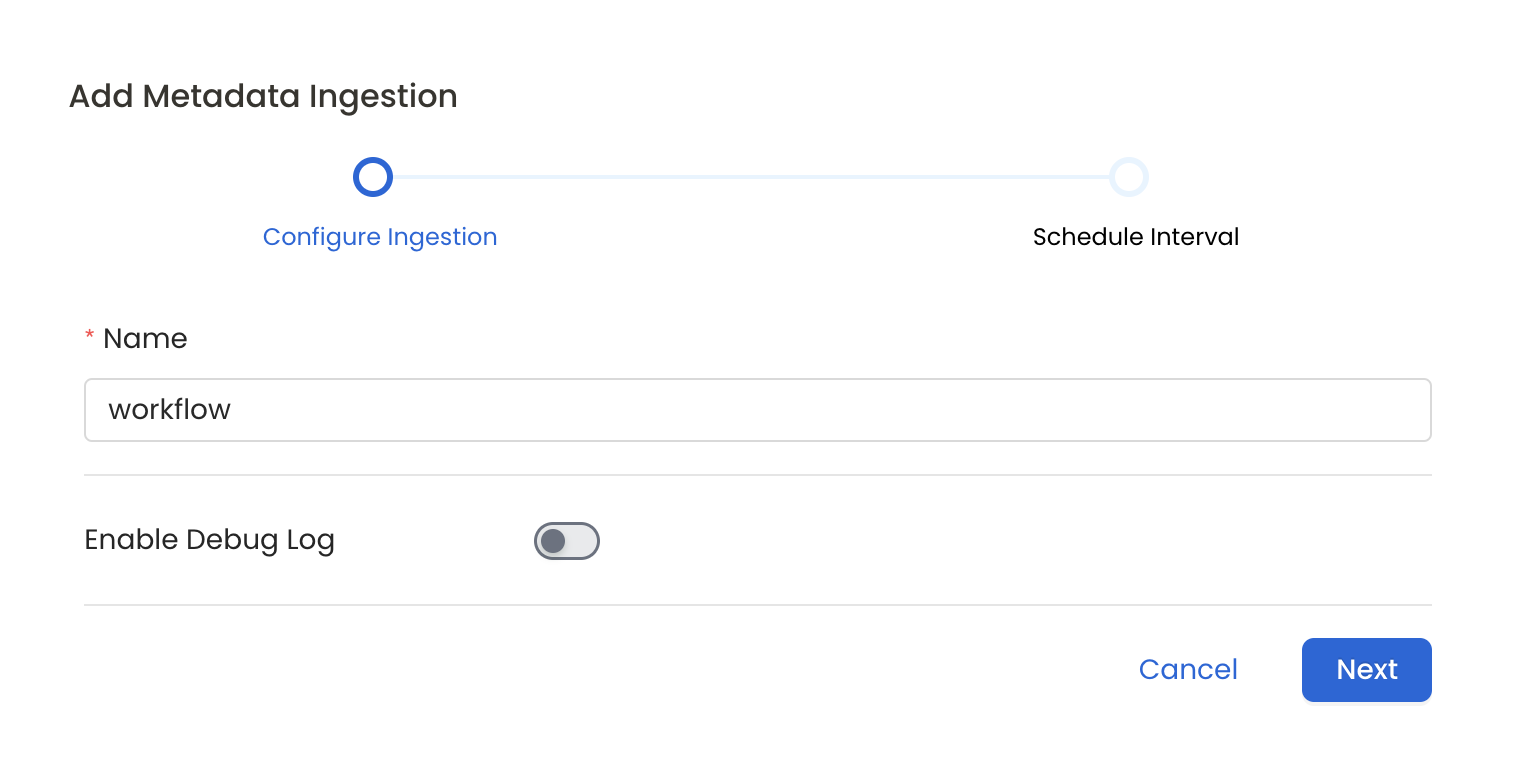

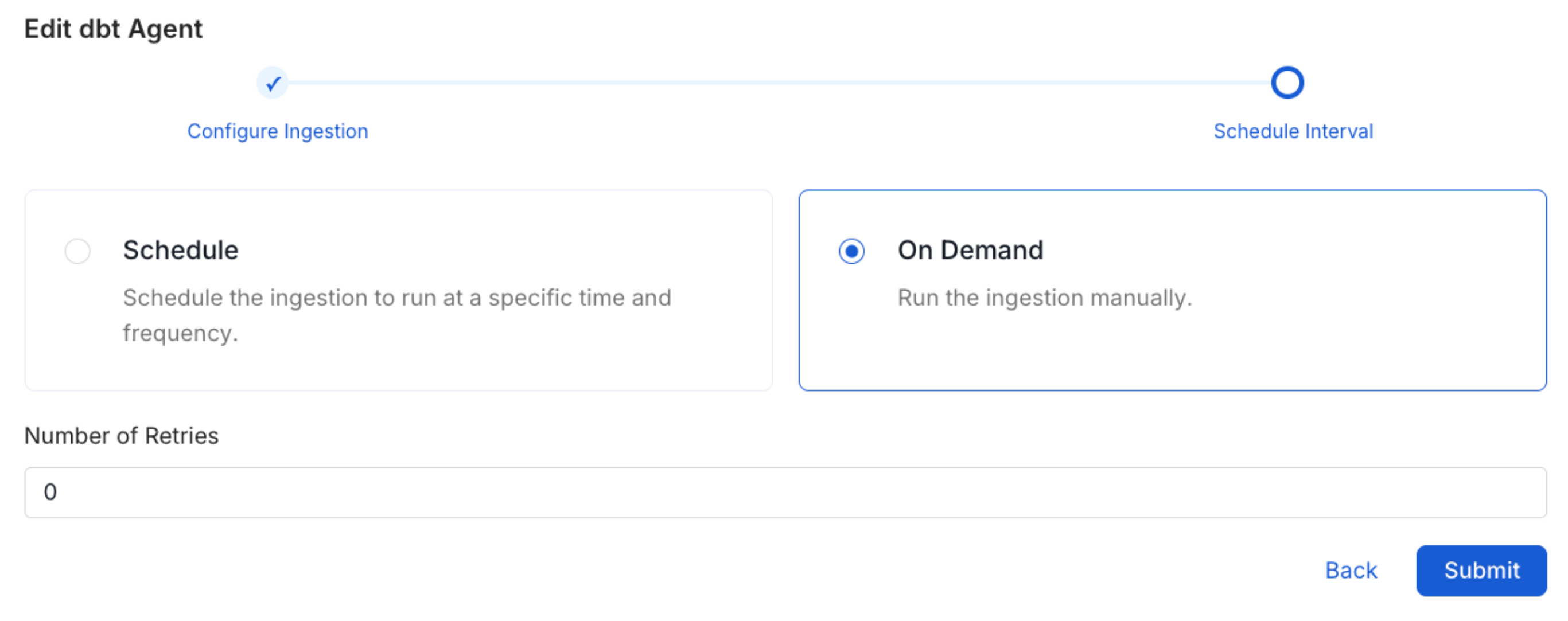
Schedule the Ingestion and Deploy
Scheduling can be set up at an hourly, daily, weekly, or manual cadence. The
timezone is in UTC. Select a Start Date to schedule for ingestion. It is
optional to add an End Date.Review your configuration settings. If they match what you intended,
click Deploy to create the service and schedule metadata ingestion.If something doesn’t look right, click the Back button to return to the
appropriate step and change the settings as needed.After configuring the workflow, you can click on Deploy to create the
pipeline.